Vmd files2/16/2024 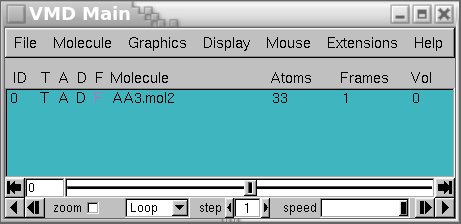
When observing molecular simulations, it is not uncommon to encounter a lack of smoothness in the visual representation. This will open up the Graphical Representations menu that you can use to customize the aspect of your system. You can select the representation that is more suitable for your specific case by going to Graphics $\rightarrow$ Representations. One of the key features of VMD is its ability to create diverse and customizable representations of molecular structures, leading to enhanced visualization and comprehension of the trajectory data.īy leveraging the representation tools in VMD, you can create visual depictions that bring clarity and coherence to your molecular simulations. You can use the buttons in this panel to reproduce the trajectory, adjust the speed to your preference, and even modify the number of steps you want to show in the GUI.Ĭreating representations for enhanced visualization Player Controls: The VMD player includes intuitive controls for playing the trajectory as a movie. This functionality facilitates focused analysis and examination of particular frames of significance. You can also use it as a convenient text entry, to specify a certain frame you are interested in. Slider: The slider is a fundamental control that enables navigation from one frame to another by simply dragging it.Ĭurrent Frame Selection: This panel tells you the frame you are currently looking at. Total Number of Frames: This panel displays the overall number of frames present in the loaded trajectory. When working with the VMD player, there are four key panels to keep in mind: The VMD player facilitates the visualization and exploration of molecular dynamics trajectories in a dynamic and interactive manner.  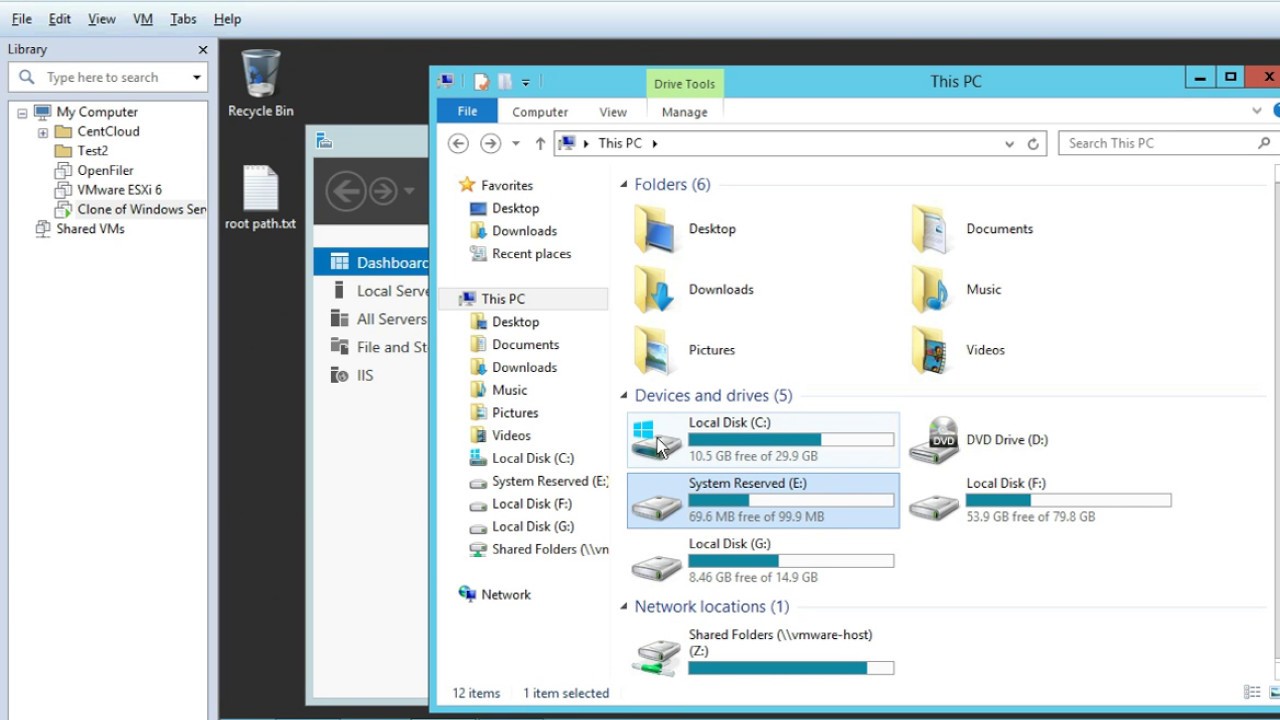
Once the trajectory is loaded into VMD, you can navigate through it using the VMD player, which is a powerful tool within the software. Visualizing and Navigating Trajectories in VMD To obtain the gro and xtc files with the same number of atoms you can use the gmx trjconv command as shown here If you try to load the entire xtc file containing also other atoms ( e.g., water) then you will not be able to do so. This is probably due to the fact that the two files provided have a non-matching number of atoms.Īlways remember that the number of atoms in the gro and xtc files needs to be the same otherwise you won’t be able to correctly load the trajectory.įor instance, if your gro file only contains the protein, then you will need to load the xtc file that only contains the velocities and coordinates of the protein. 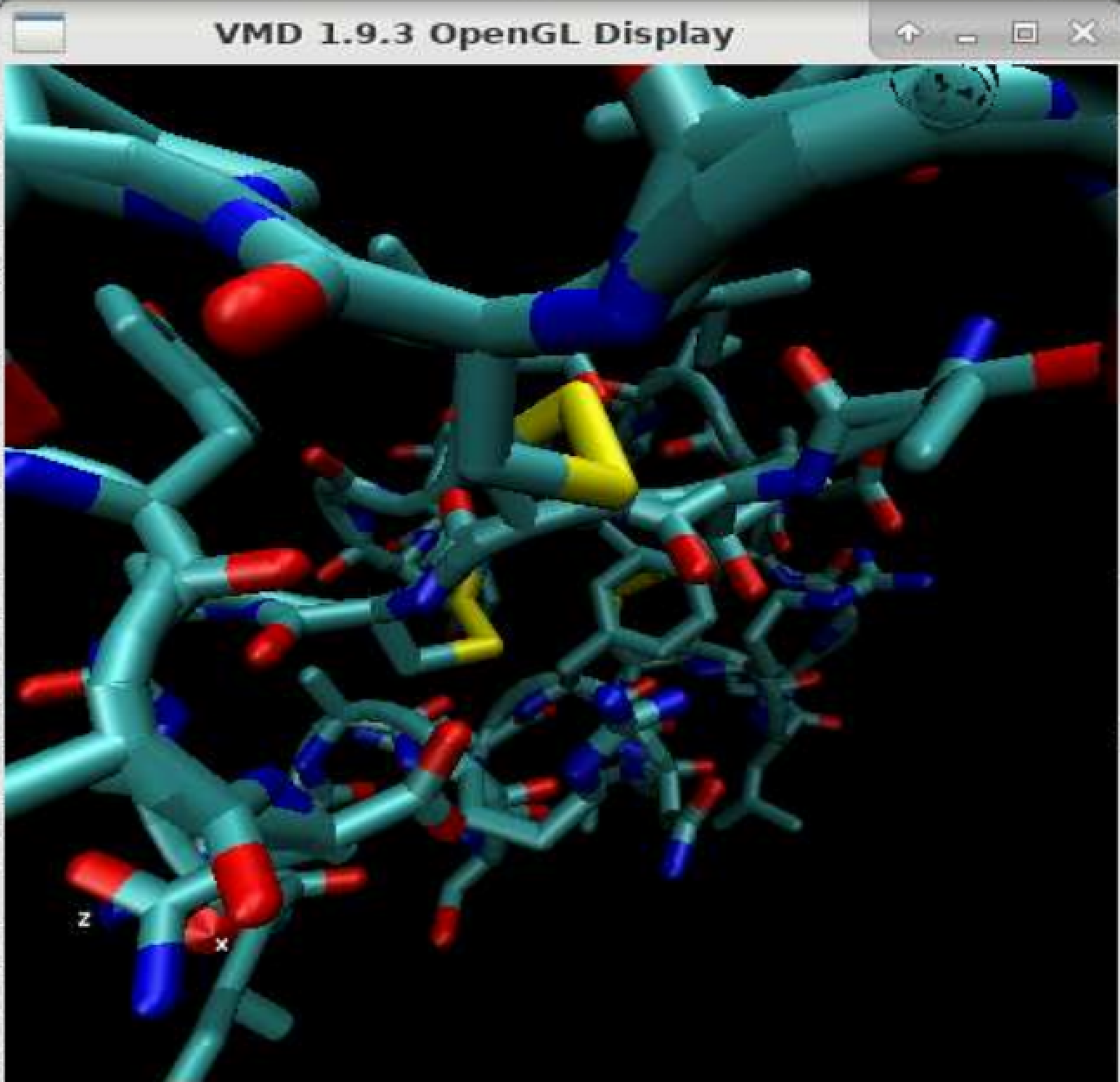
Sometimes you may encounter an error or you may notice that the trajectory is not loaded. You can simply go to the directory where the files are stored and load them with this command: The first one relies on the command line. In VMD, you have two main options for loading your GROMACS trajectory. It is good practice to use the compressed trajectory format, namely the xtc file as it is less memory-consuming and therefore easier to load and move from one workstation to another. You can use both the gro file and the pdb file format.Ī GROMACS trajectory file which contains the atomic coordinates and velocities recorded throughout your simulation. Once you are all set, you only need to locate two files on your computer:Ī reference structure containing the initial coordinates for your system. To start off, make sure you have VMD installed and up and running on your system. This is pretty straightforward once you know how to do it. Loading GROMACS trajectories in VMD: Step-by-Step GuideĪs you may imagine, the first step will be to load your trajectory in VMD. The main focus of this article is to show you how to load your xtc trajectories in VMD and then how to navigate and observe them so that later on you can extract relevant information hidden in your simulations. In this blog post, I will explain how you can achieve it using Visual Molecular Dynamics (VMD), which despite its outdated interface is still the go-to tool to visualize molecular trajectories. Ok, so you have run your molecular simulation with GROMACS, and now you would like to have a visual representation to gain a deeper understanding of the dynamical behavior of your system.
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
